DAVID BAKER SPENDS a large part of each weekday in back-to-back, conversations, dreaming up new proteins that could change the world. What if an enzyme could break down plastic into non-toxic components? Could a protein bind to virus cells and render them harmless? Can the properties of silk, abalone shell and tooth be borrowed to make powerful new eco-friendly materials?
Baker’s lab, at the University of Washington Institute for Protein Design, made an incredible breakthrough developing technology that enables scientists to design proteins never seen before in nature. Before, scientists studied and tweaked a finite number of proteins that evolved slowly over the lifespan of the planet. Now we’re entering a new era, as he said in his 2019 TED talk, in which brand-new proteins can be built from scratch.
“There’s such a wide range of possibilities. A lot of the ingenuity and creativity is about the question of what to design.”
David Baker, University of Washington Institute for Protein Design
Suddenly, scientists in the field have an extraordinary amount of freedom. Like artists, they are bound only by their imaginations. Perhaps they need to think differently, to come up with creations as ingenious as nature’s designs.
“There’s such a wide range of possibilities,” Baker says over the phone as he walks near his home in Seattle, Washington. “A lot of the ingenuity and creativity is about the question of what to design. It should be interesting, potentially useful, presumably new. That’s up to the imagination of the designer, and good ideas come from all over the place.”
One of his strategies for boosting creativity within his team is to eliminate any sense of hierarchy that could stifle innovation. “Everyone’s at the same level; everyone’s ideas are respected.” Another is to bounce ideas around constantly.
“I really believe in this communal brain concept,” he says, “which is that great ideas come from interactions between smart, energetic people who are excited about working on the same problem. Connectivity between neurons is where you get all sorts of new emergent properties and I really think that’s true of science as well. I’m just one neuron that happens to have a lot of different connections.”

Throwing together ideas generated by diverse minds and seeing what new possibilities emerge is one way to spur creativity. Another is for each person to develop a wide range of interests and experiences. Baker studied philosophy and social science before switching to biology. He says he likes to read a lot in his spare time, as well as getting out in nature when he can. This is why he’s based in the Pacific Northwest, with its mountains, forests and coastlines.
It was while he was still an undergrad at Harvard that Baker read a book on molecular biology within cells and changed his path. He undertook a postdoc with structural biologist David Agard at University of California, San Francisco, in 1989, and by 1993, when he joined the University of Washington, he had become fascinated by the topic of protein folding.
It was a problem that had always stumped scientists in the field. Proteins are the building blocks of life: molecules made up of strings of amino acids. They carry out all the major tasks in our bodies, from fending off disease to carrying messages in our blood to firing neurons. Any sequence of amino acids spontaneously fold into a complex three-dimensional structure that’s determined by multiple forces, but it was impossible to predict how a particular sequence would fold until it happened.
Then, Baker and his group started developing a software programme called Rosetta in the late 90s that had some degree of success in making these predictions. It’s open-source and has been added to collaboratively in the decades since. “Now,” Baker says, “we’re increasingly incorporating deep learning and AI into [Rosetta], and it’s definitely making it more powerful.”
AI is rapidly improving at predicting a protein’s shape, or how it will fold, based on its amino acid sequence. Ultimately, this brings scientists closer to being able to dream up brand-new designs, and then build the amino acid chain that will produce the new protein. Scientists are imagining new types of protein, from coronavirus therapeutics and cancer cures to new types of solar energy generation.

“What you’re doing with AI is accelerating what nature might do in a million years.”
Nicolas Lopez Carranza, Head of BioAI at InstaDeep
“The way protein design works is you begin with an initial protein that you want to optimise,” explains Nicolas Lopez Carranza, InstaDeep’s Head of BioAI who leds the team behind the AI protein design platform DeepChain. “You want to find the mutations that, say for example, make it bind better. What you’re doing with AI is accelerating what nature might do in a million years. But you need it now.”
For proteins, the sequence of amino acids determines the shape, and the shape determines function. “It’s all about shape,” Lopez Carranza says. “Protein design is finding the right shape to make a protein’s interactions more efficient – or less efficient – depending on the specific problem you want to solve.”
What excites him most about this new age of protein design is that breakthroughs in biology will not come exclusively from biology experts anymore. The state of the art for protein-shape prediction is based on the Critical Assessment of Protein Structure Prediction, or CASP, a contest also known as “the Olympics of protein-folding prediction.”
He points to the last two CASP competitions where “biology rookies” who were experts in AI beat labs with 30 years of experience in protein folding.
One of the most celebrated examples of protein design – so far –is Frances Arnold, a professor at the California Institute of Technology, who was awarded the Nobel Prize in Chemistry in 2018 for pioneering the use of directed evolution to engineer enzymes. AI machine learning methods have made it possible to emulate her Nobel prize-winning work mimicking natural selection by encouraging mutations, selecting for certain traits, and using the best candidates as the starting point for a new round of mutations.
“It’s a lot easier to design in silico [on a computer] than to actually do the chemistry. It means that the rate of progress will accelerate.”
-Byron Rubin, protein crystallographer-turned sculptor
Chris Voigt is another pioneer who has been dubbed “biology’s toymaker” for his knack for building protein machines. The MIT synthetic biologist, molecular biophysicist, and engineer has worked on directed-evolution projects that engineer E.coli bacteria to invade cancer cells, where they could potentially deliver targeted chemotherapy drugs.
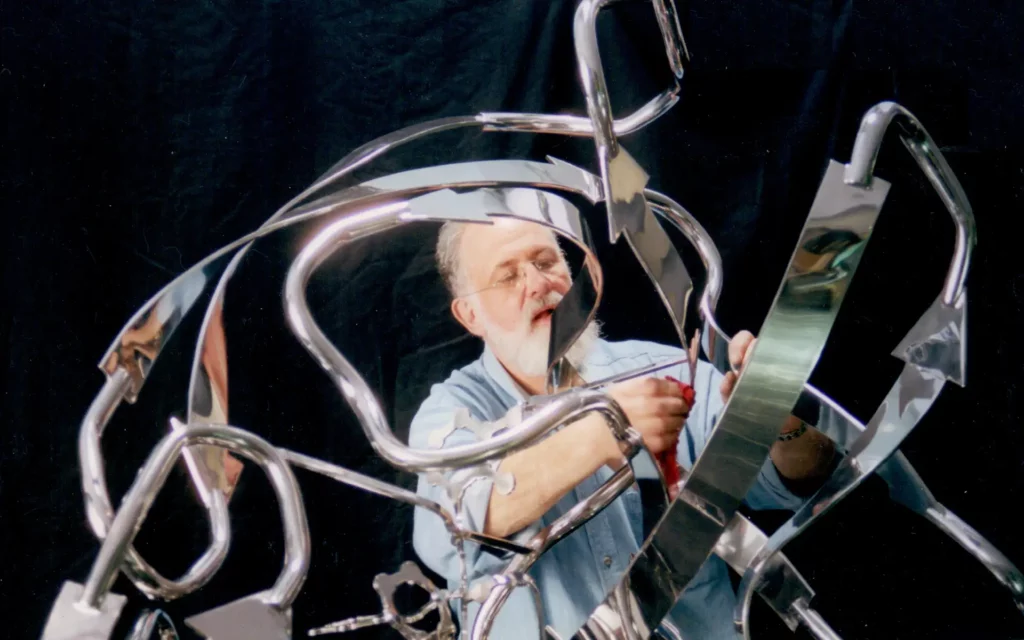
“De novo” protein design is still in its infancy, compared with these techniques, but its implications are far-reaching. “The puzzle, in many ways, is solved,” says Byron Rubin, a former crystallographer, whose job once was to determine the structure of proteins by painstakingly uncovering the arrangements of atoms within them, using x-rays. “It’s a lot easier to design in silico [on a computer],” he says “than to actually do the chemistry. It means that the rate of progress will accelerate.”
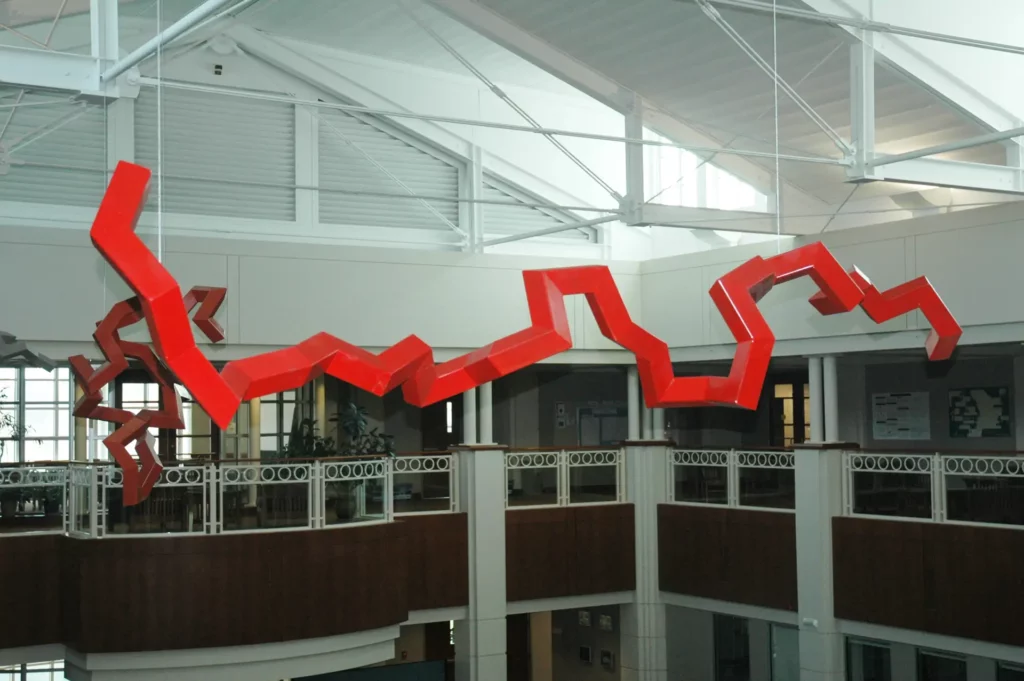
Rubin was an undergrad at Reed University in the late 60s, when the molecular structure of only a handful of proteins was known. Models of these were created using wooden dowels and plastic balls, but in 1972 he invented a wire-bending machine that allowed them to be depicted in a new way, with the ‘backbone’ of each structure twisting, folding and spiralling. By the 1990s, Rubin had switched to sculpting full-time, and if you now visit Pfizer’s research campus in La Jolla, California, you’ll see an eight-foot stainless steel sculpture of an anti-HIV drug’s molecular structure in the lobby that he made. It’s one of many monumental works of art he has built that are based on protein molecules.
“There’s a lot of artists doing science, let me tell you,” Rubin says from his home in Rochester, New York. “There always has been. Taking disparate data and assembling it into a coherent theory, that’s an art in itself.”
While many artists are inspired by science, scientists can also learn a lot from the realm of art: at least that’s what DePauw University biochemistry professor Daniel Gurnon believes. In 2011, he invited another scientist-turned-sculptor, Julian Voss-Andreae, to help his students weld protein sculptures to gain a better understanding of the field.
“A big part of science, particularly biochemistry,” he says, “involves using artistic license to represent concepts that our human minds did not evolve to observe directly. I’m talking about interactions on timescales much faster or slower than we can conceive, or wavelengths of light that are not just invisible but unseeable.”
Look up what a protein structure is now and you’ll see “beautiful twisting ribbons and coils,” he adds, but this is “an artistic rendition that reveals just the important bits. There are different ways of depicting the truth. I felt pretty strongly that science has an important connection to the arts. Imagining the world of molecules takes a lot of creativity.”

Voss-Andreae was embedded with a team of Caltech and NYU scientists several years ago who were creating new nanomaterials by rearranging DNA, and he, too, sees important similarities between the worlds of art and science. What can be helpful in both cases, he says, is “having a mindset that you are essentially playing. You’re not only in control, figuring it all out, you’re playing with a system and it will all fall into place. History’s full of that stuff.”
Baker’s lab recognises the need for play in the serious business of science. In 2008, the team launched a videogame-like interface for the Rosetta software called Foldit, and allowed anyone to try their hand at designing new proteins. The idea, says research scientist Brian Koepnick, who heads up the initiative, was to crowdsource the problem of protein structure prediction, and eventually protein design itself.
“We recognized the value of Foldit players’ creativity,” he says. “If you ask 100 Foldit players to solve a problem, they will try 100 different approaches.” A trained scientist will most likely create something that looks similar to a natural protein, Koepnick adds. But “Foldit players don’t have these preconceived notions about protein structures, so they will design crazy new protein folds that are completely unlike natural proteins.” It’s this “lateral thinking,” he believes, “that is the most exciting aspect of Foldit creativity.”
That intuitive play of twisting proteins into particular shapes caught the attention of Demis Hassabis, the co-founder of DeepMind.
“Somehow these gamers had trained their intuition and their pattern-matching capabilities so that somehow they were able to do what brute-force computer systems couldn’t at the time – and actually come up with the right shapes,” he told the BBC.
“That made me think that AI could maybe try to mimic that intuitive capability that those gamers were demonstrating.”
The result was AlphaFold, a revolutionary AI breakthrough for predicting protein folding that won the last two CASP competitions.
Computing power is an indispensable part of the protein-design revolution, but human creativity still plays an important role. Even beauty can play a part in scientific progress: both Albert Einstein and Paul Dirac, who predicted antimatter and was awarded a shared Nobel Prize in 1933, talk about aesthetic value in relation to their work. Dirac “said there must be an antiparticle for the electron, because it’s just beautiful that way,” Voss-Andreae says. “And it turned out to be true.”
Aesthetics are only one part of the story, of course. As much as new-generation protein designers are artists, they are also engineers, working backwards from a desired function, and creating a structure that fulfils this purpose. Nature doesn’t always have a great sense of style, Voss-Andreae has found. He tends to build sculptures depicting molecules he finds both visually and conceptually attractive, such as collagen and haemoglobin, but there aren’t many of those.
“There are weird little viruses that I love,” he says. But COVID-19, for example, “is an ugly little guy. The spikes on the outer shell don’t appear to be regular, which looks strange to me. Corona, AIDS, I don’t like them so much.”
Then again, art isn’t always about beauty either. For every Michaelangelo, creating harmony and balance, there is a successful Jackson Pollock or Marcel Duchamp, making us reconsider what we find valuable. Sometimes thinking like an artist doesn’t mean making something pretty; sometimes it means breaking all the rules, and making something that’s never been seen before.




